JMJD6 (Gene)
Analyze
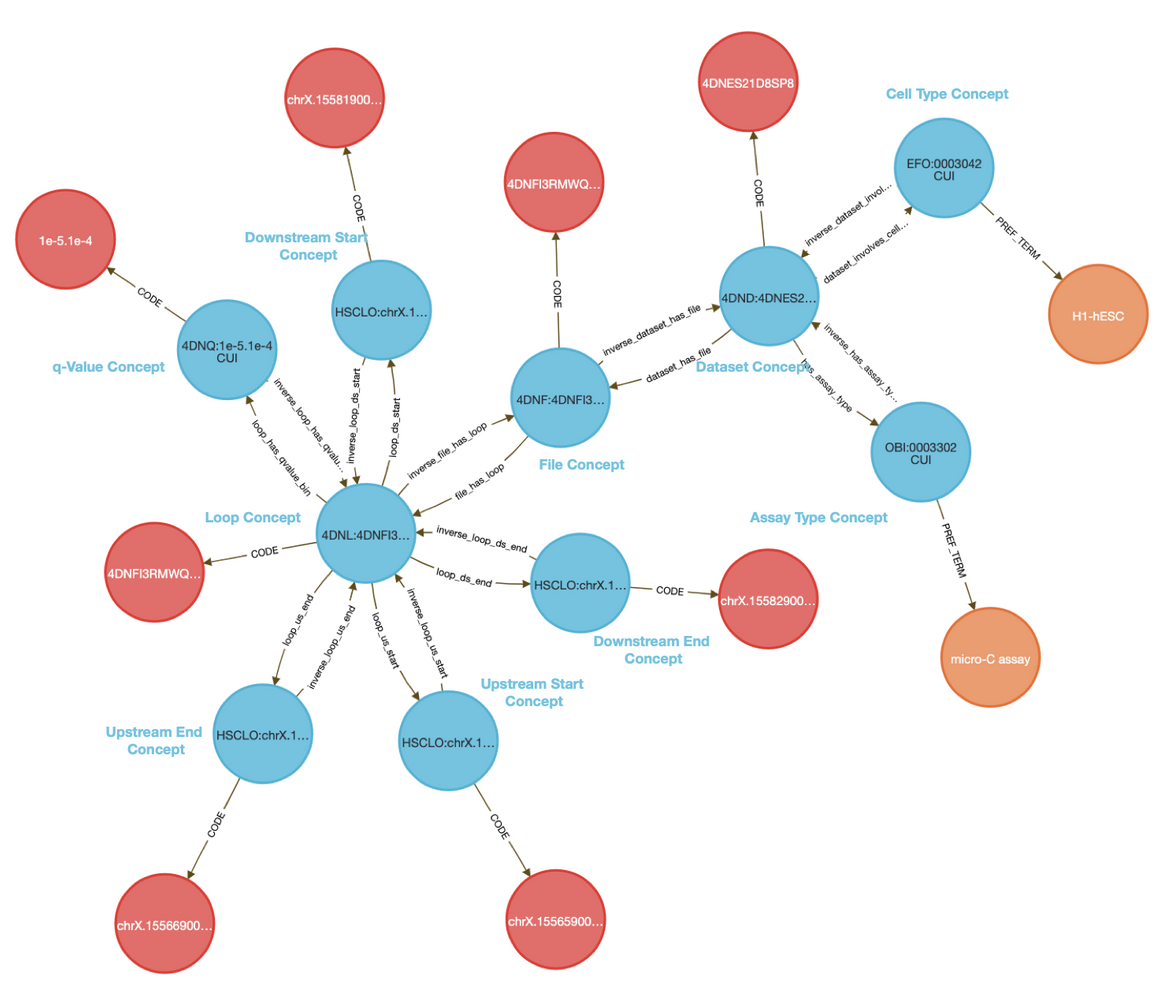
The CFDE Data Distillery Knowledge Graph contains entities and relationships across the CFDE. View JMJD6's neighborhood in the knowledge graph.
View Gene-Centric information about the gene from a pre-built PWB workflow. View the workflow with JMJD6.

The CFDE Gene Centric Appyter Resolves and Displays Gene-Centric information from CFDE APIs. Execute the Appyter using JMJD6.

The Gene and Drug Landing Page Aggregator (GDLPA) finds links to primary and secondary source information from CFDE and other resources. Discover landing pages for JMJD6.

The Playbook Workflow Builder helps you interactively construct workflows leveraging CFDE APIs without code. Start a new workflow with JMJD6.
Results found
Linked to
| Label | Description |
|
|---|---|---|---|
A Gene Set from LINCS | |||
A Gene Set from LINCS | |||
A Gene Set from LINCS | |||
A Gene Set from LINCS | |||
A Gene Set from LINCS | |||
A Gene Set from LINCS | |||
A Gene Set from LINCS | |||
A Gene Set from LINCS | |||
A Gene Set from LINCS | |||
A Gene Set from LINCS |
A Gene Set from LINCS
A Gene Set from LINCS
A Gene Set from LINCS
A Gene Set from LINCS
A Gene Set from LINCS
A Gene Set from LINCS
A Gene Set from LINCS
A Gene Set from LINCS
