H1-2 (Gene)
Analyze
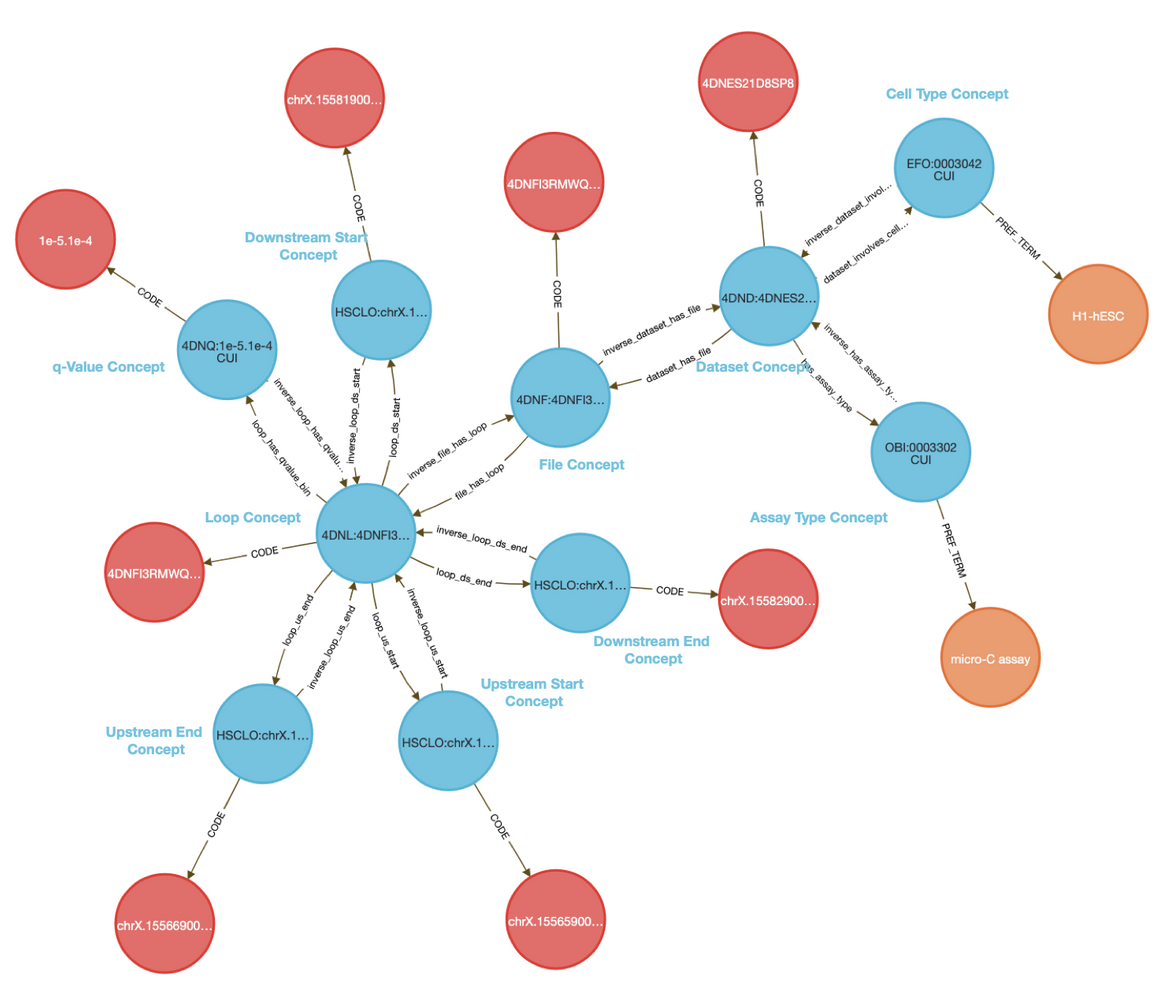
The CFDE Data Distillery Knowledge Graph contains entities and relationships across the CFDE. View H1-2's neighborhood in the knowledge graph.
View Gene-Centric information about the gene from a pre-built PWB workflow. View the workflow with H1-2.

The CFDE Gene Centric Appyter Resolves and Displays Gene-Centric information from CFDE APIs. Execute the Appyter using H1-2.

The Gene and Drug Landing Page Aggregator (GDLPA) finds links to primary and secondary source information from CFDE and other resources. Discover landing pages for H1-2.

The Playbook Workflow Builder helps you interactively construct workflows leveraging CFDE APIs without code. Start a new workflow with H1-2.
Results found
Linked to
| Label | Description |
|
|---|---|---|---|
A Gene Set from LINCS | |||
A Gene Set from LINCS | |||
A Gene Set from LINCS | |||
A Gene Set from LINCS | |||
A Gene Set from LINCS | |||
A Gene Set from LINCS | |||
A Gene Set from LINCS | |||
A Gene Set from LINCS | |||
Cerebellar ataxia refers to ataxia due to dysfunction of the cerebellum. This causes a variety of el... | |||
A Gene Set from LINCS |
A Gene Set from LINCS
A Gene Set from LINCS
A Gene Set from LINCS
A Gene Set from LINCS
A Gene Set from LINCS
A Gene Set from LINCS
A Gene Set from LINCS
A Gene Set from LINCS
Cerebellar ataxia refers to ataxia due to dysfunction of the cerebellum. This causes a variety of el...
A Gene Set from LINCS

